Statistical Data Analysis |
Next Generation Sequencing |
Genomics & Proteomics
Microarray data analysis
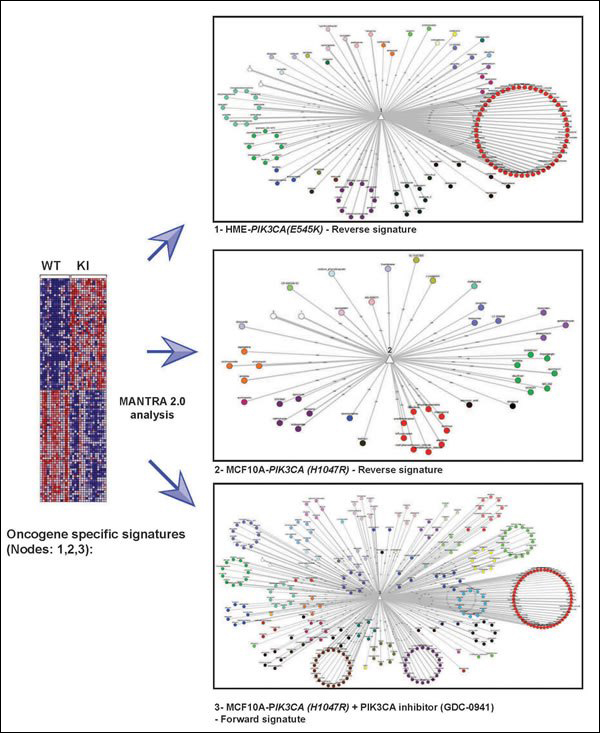
Microarray analysis techniques are used in interpreting the data generated from experiments on DNA, RNA, and protein microarrays, which allow researchers to investigate the expression state of a large number of genes - in many cases, an organism's entire genome - in a single experiment.
Microarray analysis: reference
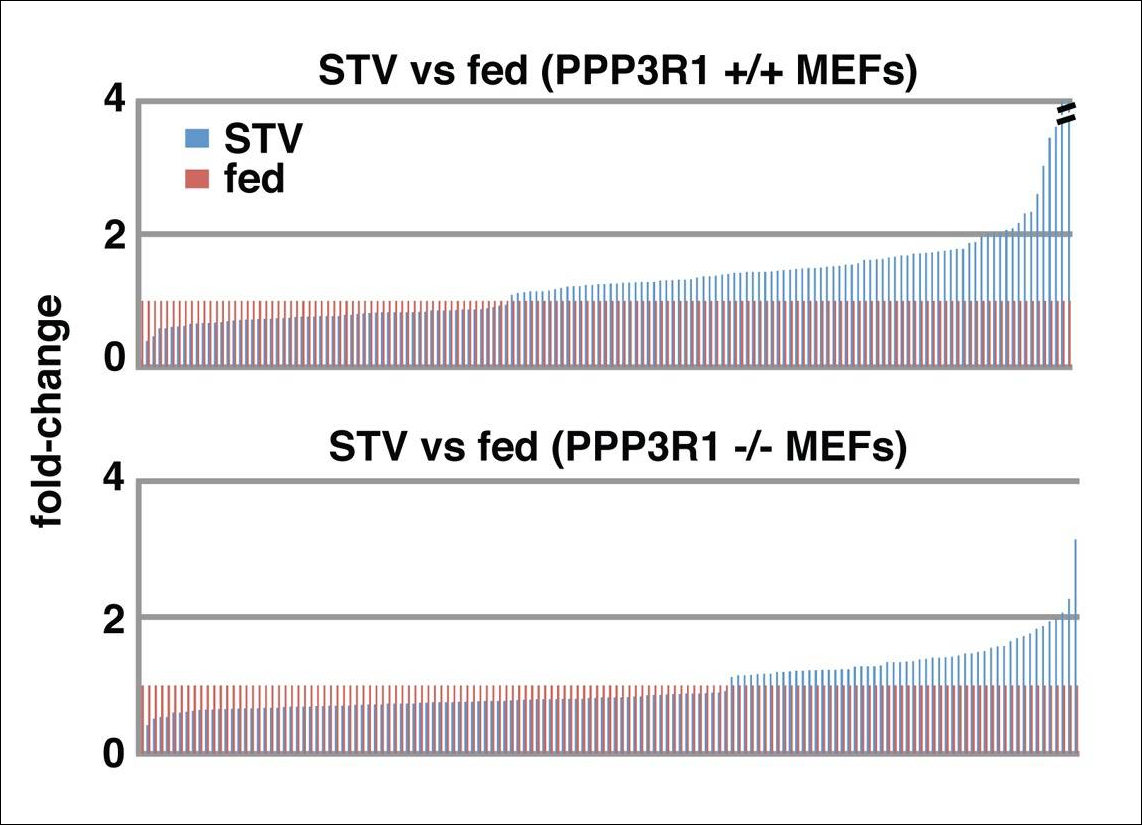
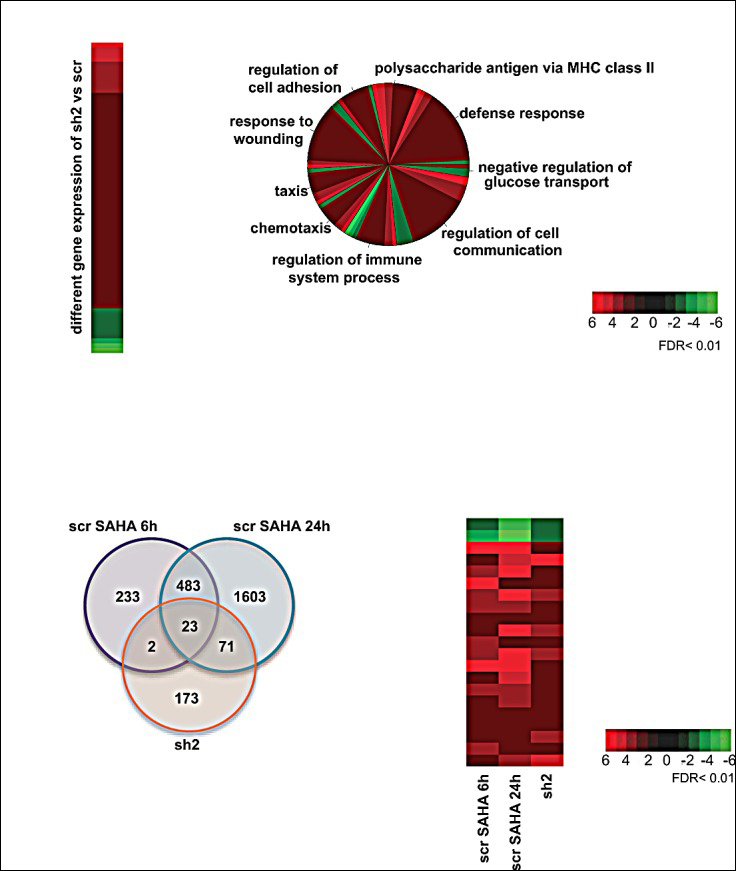
Transcriptional response: reference
Submission of data in NCBIs Gene Expression Omnibus (GEO)
A fundamental principle guiding the publication of scientific results is that the data supporting any scholarly work must be made fully available to the research community, in a form that allows the basic conclusions to be evaluated independently. In the context of molecular biology, this has typically meant that authors of a paper describing a newly sequenced genome, gene, or protein must deposit the primary data in a permanent, public data repository, such as the sequence databases maintained by the National Center for Biotechnology Information (NCBI).
Gene Networks
A gene network is a collection of molecular regulators that interact with each other and with other substances in the cell to govern the gene expression levels of mRNA and proteins.
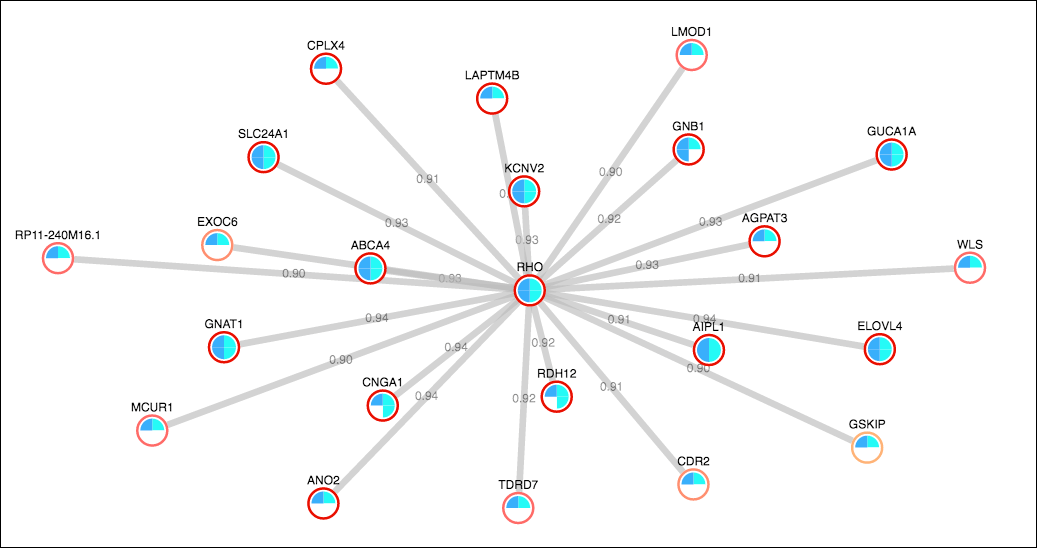
Gene Network: reference
Pathway analysis
Gene Set Enrichment Analysis and Gene Ontology analysis (meta-analysis to infer biological function for unknown genes).
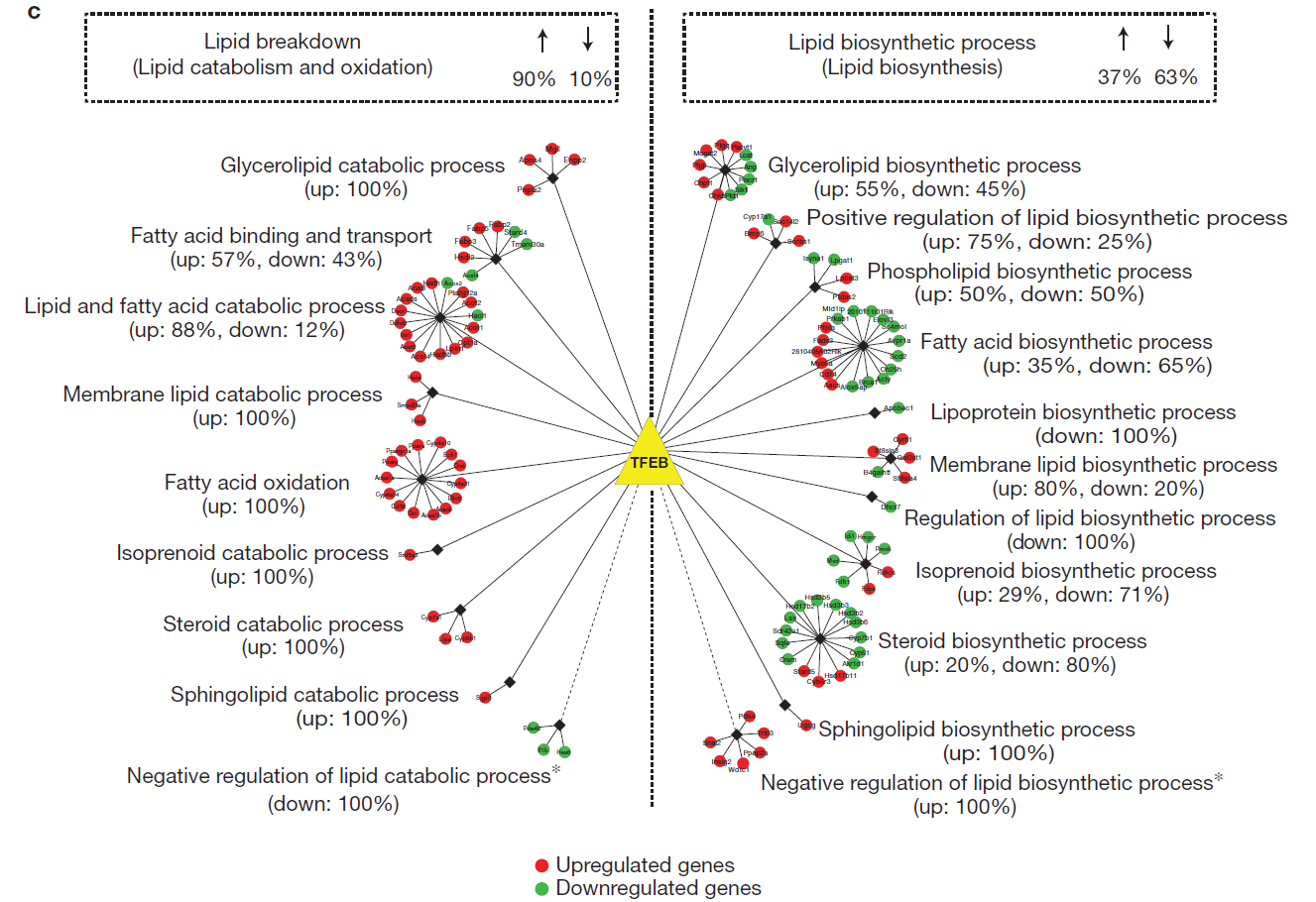
Pathway Network: reference
Transcription Factor Binding Site
Identification of putative transcription factor binding sites (TFBS) in promoter gene sequences from a species or more species of interest.
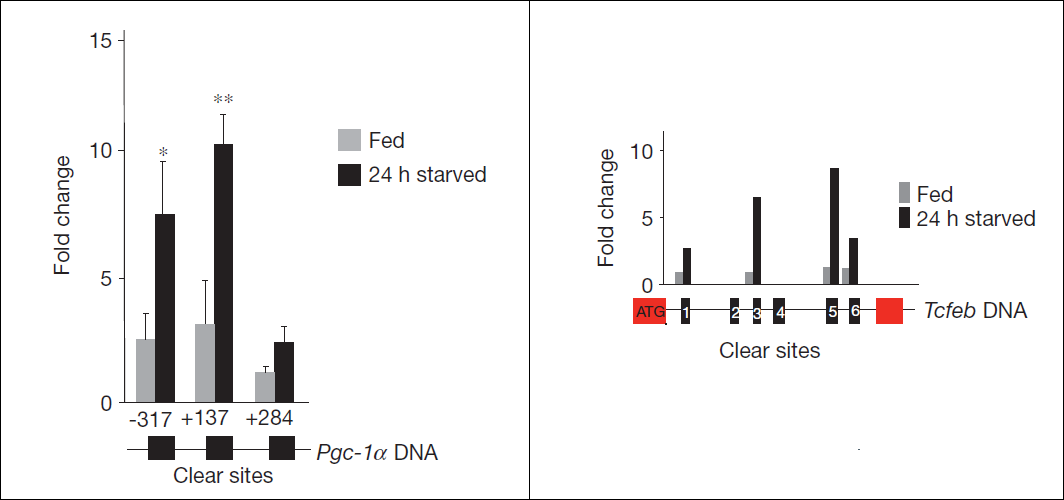
Binding Sites: reference
Gene annotation
Gene identification from sequences, gene identifiers conversion, gene characterization, sequence feature (exon, intron, ORF, CDS) identification
Integration of data results from different sources
Data integration involves combining data residing in different sources and providing users with a unified view of these data. This process becomes significant in scientific domains, combining research results from different bioinformatics repositories.
Protein analysis
Protein domain analysis, Protein-protein interaction, Protein localization prediction.
Drug Networks
A drug network is a collection of drugs and treatments correlated to each other according to transcriptional or structural similarity.